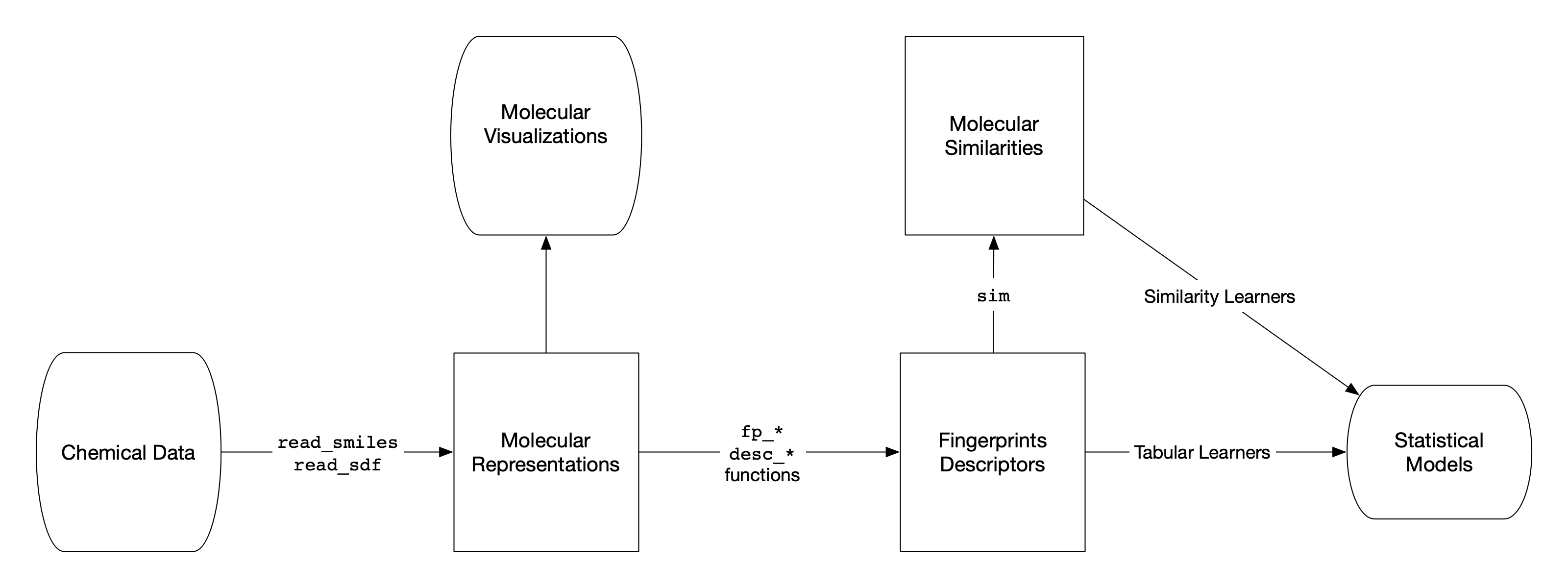
Read, preprocess, and featurize chemical data using RDKit.
A lightweight R interface for accessing RDKit via the RDKit Python API.
Install rdkit
A simple way to install the Python package rdkit that tidychem can discover easily, is to run the helper function tidychem::install_rdkit(). By default, it installs rdkit into an virtual environment named r-rdkit.
tidychem::install_rdkit()This follows the best practices suggested by the reticulate vignette Managing an R Package’s Python Dependencies. There are also recommendations in the vignette on how to manage multiple R packages with different Python dependencies.
Usage
library("tidychem")
"smi-multiple.smi" |>
tidychem_example() |>
read_smiles() |>
fp_morgan()